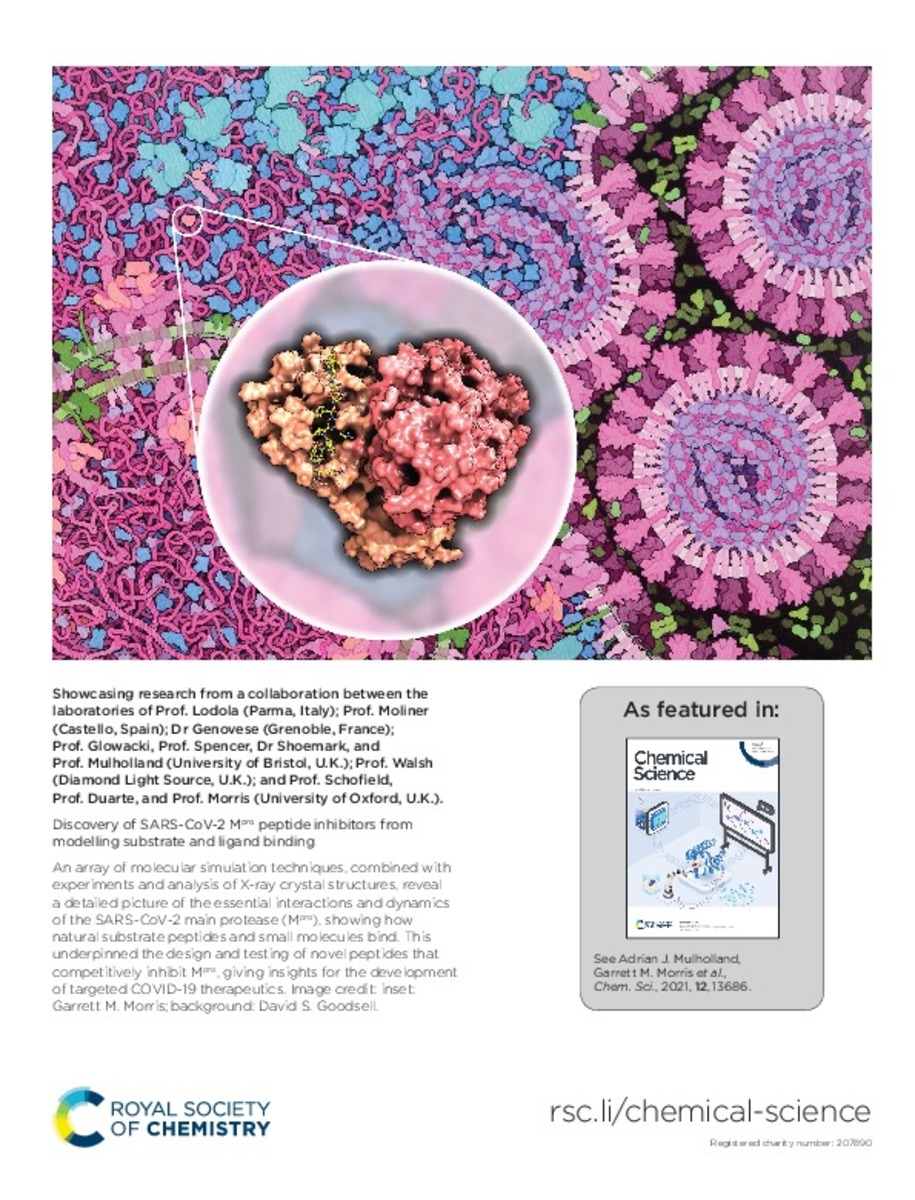
Discovery of SARS-CoV-2 M-pro peptide inhibitors from modelling substrate and ligand binding
Ver/
Impacto
 Scholar |
Otros documentos de la autoría: Chan, Henry; Moesser, Marc Alexander; Walters, Rebecca; Malla, Tika; Twidale, Rebecca; John, Tobias; Deeks, Helen M.; Johnston-Wood, Tristan; Mikhailov, Victor; Sessions, Richard; Dawson, William; Salah, Eidarus; Lukacik, Petra; Strain-Damerell, Claire; Owen, David; Nakajima, Takahito; Świderek, Katarzyna; Lodola, Alessio; Moliner, Vicent; Glowacki, David; Walsh, Martin Austin; Schofield, Christopher; Genovese, Luigi; Shoemark, Deborah K.; Mulholland, Adrian; Duarte, Fernanda; Morris, Garrett
Scholar |
Otros documentos de la autoría: Chan, Henry; Moesser, Marc Alexander; Walters, Rebecca; Malla, Tika; Twidale, Rebecca; John, Tobias; Deeks, Helen M.; Johnston-Wood, Tristan; Mikhailov, Victor; Sessions, Richard; Dawson, William; Salah, Eidarus; Lukacik, Petra; Strain-Damerell, Claire; Owen, David; Nakajima, Takahito; Świderek, Katarzyna; Lodola, Alessio; Moliner, Vicent; Glowacki, David; Walsh, Martin Austin; Schofield, Christopher; Genovese, Luigi; Shoemark, Deborah K.; Mulholland, Adrian; Duarte, Fernanda; Morris, Garrett
Metadatos
Mostrar el registro completo del ítemcomunitat-uji-handle:10234/9
comunitat-uji-handle2:10234/160292
comunitat-uji-handle3:10234/160293
comunitat-uji-handle4:
INVESTIGACIONMetadatos
Título
Discovery of SARS-CoV-2 M-pro peptide inhibitors from modelling substrate and ligand bindingAutoría
Fecha de publicación
2021-09-06Editor
The Royal Society of ChemistryCita bibliográfica
CHAN, HT Henry, et al. Discovery of SARS-CoV-2 Mpro Peptide Inhibitors from Modelling Substrate and Ligand Binding. bioRxiv, 2021. doi: https://doi.org/10.1101/2021.06.18.446355Tipo de documento
info:eu-repo/semantics/articleVersión
info:eu-repo/semantics/publishedVersionPalabras clave / Materias
Resumen
The main protease (Mpro) of SARS-CoV-2 is central to viral maturation and is a promising drug target, but
little is known about structural aspects of how it binds to its 11 natural cleavage sites. We used
biophysical ... [+]
The main protease (Mpro) of SARS-CoV-2 is central to viral maturation and is a promising drug target, but
little is known about structural aspects of how it binds to its 11 natural cleavage sites. We used
biophysical and crystallographic data and an array of biomolecular simulation techniques, including
automated docking, molecular dynamics (MD) and interactive MD in virtual reality, QM/MM, and linearscaling DFT, to investigate the molecular features underlying recognition of the natural Mpro substrates.
We extensively analysed the subsite interactions of modelled 11-residue cleavage site peptides,
crystallographic ligands, and docked COVID Moonshot-designed covalent inhibitors. Our modelling
studies reveal remarkable consistency in the hydrogen bonding patterns of the natural Mpro substrates,
particularly on the N-terminal side of the scissile bond. They highlight the critical role of interactions
beyond the immediate active site in recognition and catalysis, in particular plasticity at the S2 site.
Building on our initial Mpro-substrate models, we used predictive saturation variation scanning (PreSaVS)
to design peptides with improved affinity. Non-denaturing mass spectrometry and other biophysical
analyses confirm these new and effective ‘peptibitors’ inhibit Mpro competitively. Our combined results
provide new insights and highlight opportunities for the development of Mpro inhibitors as anti-COVID-
19 drugs. [-]
Publicado en
Chem. Sci., 2021, 12, 13686Entidad financiadora
EPSRC Centre for Doctoral Training in Synthesis for Biology and Medicine | EPSRC University of Oxford Mathematics, Physical, and Life Sciences Division (MPLS) Doctoral Training Partnership (DTP) | GlaxoSmithKline | Biotechnology and Biological Sciences Research Council (BBSRC) | Royal Society of Chemistry | BrisSynBio | EPSRC Synthetic Biology Research Centre | British Society for Antimicrobial Chemotherapy | Barcelona Supercomputing Center | UK High-End Computing Consortium for Biomolecular Simulation | Advanced Computing Research Centre, University of Bristol | Oracle Public Cloud Infrastructure | HECBioSim, ARCHER/ARCHER2 | RIKEN, HPCI System Research Project | Very Large Computing center of CEA (TGCC) | MaX Center of Excellence | Wellcome Trust, Cancer Research, UK
Código del proyecto o subvención
EP/ L015838/1 | EP/R513295/1 | BB/M011224/1 | EP/L015722/1 | EP/R512060/1 | URF\R\180033 | EP/M022609/1 | EP/N013573/1 | BB/L01386X/1 | BSAC-COVID-30 | QSB-2021-1-0007 | EP/S024093/1 | EP/L016044/1 | EP/ P020275/1 | hp200179 | hp210011 | gch0429 | gen12047 | 106244/Z/14/Z
Derechos de acceso
© 2021 The Author(s). Published by the Royal Society of Chemistry
info:eu-repo/semantics/openAccess
info:eu-repo/semantics/openAccess
Aparece en las colecciones
- INAM_Articles [506]












