QM/MM Calculations Suggest a Novel Intermediate Following the Proton Abstraction Catalyzed by Thymidylate Synthase

View/
Impact
 Scholar |
Other documents of the author: Wang, Zhen; Ferrer Castillo, Silvia; Moliner, Vicent; Kohen, Amnon
Scholar |
Other documents of the author: Wang, Zhen; Ferrer Castillo, Silvia; Moliner, Vicent; Kohen, Amnon
Metadata
Show full item recordcomunitat-uji-handle:10234/9
comunitat-uji-handle2:10234/7013
comunitat-uji-handle3:10234/8638
comunitat-uji-handle4:
INVESTIGACIONMetadata
Title
QM/MM Calculations Suggest a Novel Intermediate Following the Proton Abstraction Catalyzed by Thymidylate SynthaseDate
2013-03-07Publisher
American Chemical SocietyISSN
0006-2960; 1520-4995Bibliographic citation
WANG, Zhen, et al. QM/MM Calculations Suggest a Novel Intermediate Following the Proton Abstraction Catalyzed by Thymidylate Synthase. Biochemistry, 2013, vol. 52, no 13, p. 2348-2358Type
info:eu-repo/semantics/articlePublisher version
http://pubs.acs.org/doi/full/10.1021/bi400267qVersion
info:eu-repo/semantics/publishedVersionSubject
Abstract
The cleavage of covalent C–H bonds is one of the most energetically demanding, yet biologically essential, chemical transformations. Two C–H bond cleavages are involved in the reaction catalyzed by thymidylate synthase ... [+]
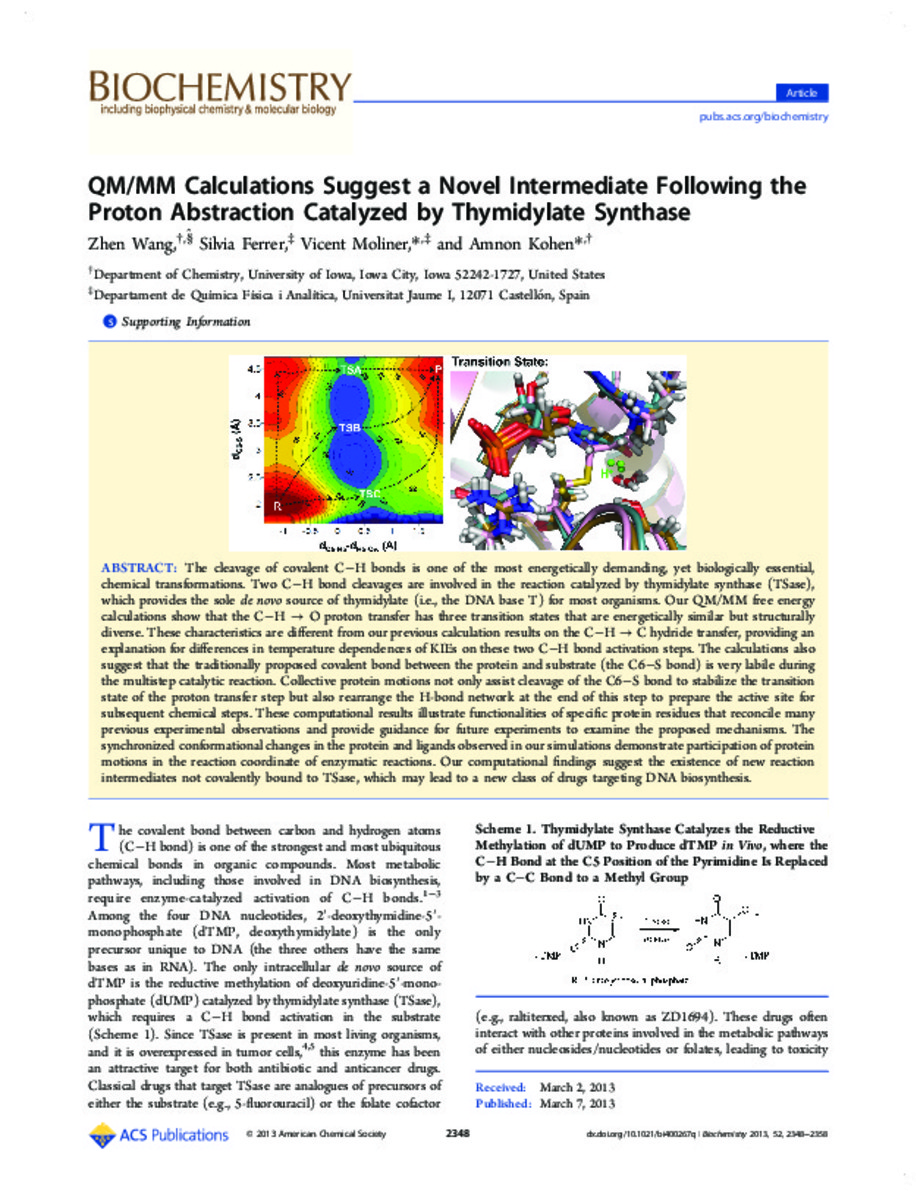
The cleavage of covalent C–H bonds is one of the most energetically demanding, yet biologically essential, chemical transformations. Two C–H bond cleavages are involved in the reaction catalyzed by thymidylate synthase (TSase), which provides the sole de novo source of thymidylate (i.e., the DNA base T) for most organisms. Our QM/MM free energy calculations show that the C–H → O proton transfer has three transition states that are energetically similar but structurally diverse. These characteristics are different from our previous calculation results on the C–H → C hydride transfer, providing an explanation for differences in temperature dependences of KIEs on these two C–H bond activation steps. The calculations also suggest that the traditionally proposed covalent bond between the protein and substrate (the C6–S bond) is very labile during the multistep catalytic reaction. Collective protein motions not only assist cleavage of the C6–S bond to stabilize the transition state of the proton transfer step but also rearrange the H-bond network at the end of this step to prepare the active site for subsequent chemical steps. These computational results illustrate functionalities of specific protein residues that reconcile many previous experimental observations and provide guidance for future experiments to examine the proposed mechanisms. The synchronized conformational changes in the protein and ligands observed in our simulations demonstrate participation of protein motions in the reaction coordinate of enzymatic reactions. Our computational findings suggest the existence of new reaction intermediates not covalently bound to TSase, which may lead to a new class of drugs targeting DNA biosynthesis. [-]
Is part of
Biochemistry, 2013, Vol. 52, nº 13Rights
Copyright © 2013 American Chemical Society
http://rightsstatements.org/vocab/InC/1.0/
info:eu-repo/semantics/openAccess
http://rightsstatements.org/vocab/InC/1.0/
info:eu-repo/semantics/openAccess
This item appears in the folowing collection(s)
- QFA_Articles [813]









