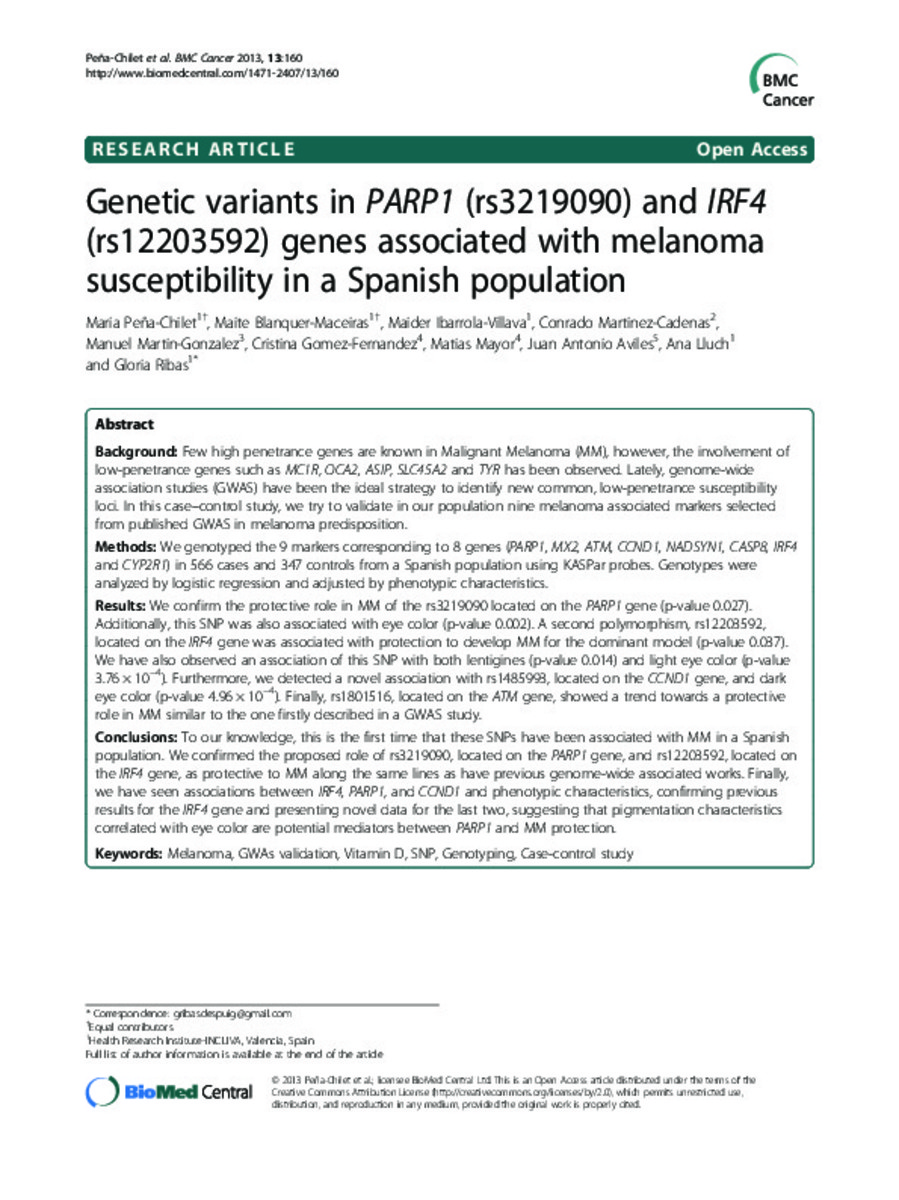
Genetic variants in PARP1 (rs3219090) and IRF4 (rs12203592) genes associated with melanoma susceptibility in a Spanish population

Ver/
Impacto
 Scholar |
Otros documentos de la autoría: Peña Chilet, María; Blanquer Maceiras, Maite; Ibarrola-Villava, Maider; Martinez-Cadenas, Conrado; Martín González, Manuel; Gómez Fernández, Cristina; Mayor, Matías; Avilés, Juan Antonio; Lluch , Ana; Ribas, Gloria
Scholar |
Otros documentos de la autoría: Peña Chilet, María; Blanquer Maceiras, Maite; Ibarrola-Villava, Maider; Martinez-Cadenas, Conrado; Martín González, Manuel; Gómez Fernández, Cristina; Mayor, Matías; Avilés, Juan Antonio; Lluch , Ana; Ribas, Gloria
Metadatos
Mostrar el registro completo del ítemcomunitat-uji-handle:10234/9
comunitat-uji-handle2:10234/36080
comunitat-uji-handle3:10234/36082
comunitat-uji-handle4:
INVESTIGACIONMetadatos
Título
Genetic variants in PARP1 (rs3219090) and IRF4 (rs12203592) genes associated with melanoma susceptibility in a Spanish populationAutoría
Fecha de publicación
2013-03Editor
BioMed CentralISSN
1471-2407; 1471-2407Cita bibliográfica
PEÑA-CHILET, Maria, et al. Genetic variants in PARP1 (rs3219090) and IRF4 (rs12203592) genes associated with melanoma susceptibility in a Spanish population. BMC cancer, 2013, vol. 13, no 1, p. 160Tipo de documento
info:eu-repo/semantics/publishedVersionVersión
info:eu-repo/semantics/publishedVersionPalabras clave / Materias
Resumen
Background
Few high penetrance genes are known in Malignant Melanoma (MM), however, the involvement of low-penetrance genes such as MC1R, OCA2, ASIP, SLC45A2 and TYR has been observed. Lately, genome-wide association ... [+]
Background
Few high penetrance genes are known in Malignant Melanoma (MM), however, the involvement of low-penetrance genes such as MC1R, OCA2, ASIP, SLC45A2 and TYR has been observed. Lately, genome-wide association studies (GWAS) have been the ideal strategy to identify new common, low-penetrance susceptibility loci. In this case–control study, we try to validate in our population nine melanoma associated markers selected from published GWAS in melanoma predisposition.
Methods
We genotyped the 9 markers corresponding to 8 genes (PARP1, MX2, ATM, CCND1, NADSYN1, CASP8, IRF4 and CYP2R1) in 566 cases and 347 controls from a Spanish population using KASPar probes. Genotypes were analyzed by logistic regression and adjusted by phenotypic characteristics.
Results
We confirm the protective role in MM of the rs3219090 located on the PARP1 gene (p-value 0.027). Additionally, this SNP was also associated with eye color (p-value 0.002). A second polymorphism, rs12203592, located on the IRF4 gene was associated with protection to develop MM for the dominant model (p-value 0.037). We have also observed an association of this SNP with both lentigines (p-value 0.014) and light eye color (p-value 3.76 × 10-4). Furthermore, we detected a novel association with rs1485993, located on the CCND1 gene, and dark eye color (p-value 4.96 × 10-4). Finally, rs1801516, located on the ATM gene, showed a trend towards a protective role in MM similar to the one firstly described in a GWAS study.
Conclusions
To our knowledge, this is the first time that these SNPs have been associated with MM in a Spanish population. We confirmed the proposed role of rs3219090, located on the PARP1 gene, and rs12203592, located on the IRF4 gene, as protective to MM along the same lines as have previous genome-wide associated works. Finally, we have seen associations between IRF4, PARP1, and CCND1 and phenotypic characteristics, confirming previous results for the IRF4 gene and presenting novel data for the last two, suggesting that pigmentation characteristics correlated with eye color are potential mediators between PARP1 and MM protection. [-]
Publicado en
BMC Cancer, 2013, vol. 13, no. 1Derechos de acceso
info:eu-repo/semantics/openAccess
Aparece en las colecciones
- MED_Articles [637]
El ítem tiene asociados los siguientes ficheros de licencia:
Excepto si se señala otra cosa, la licencia del ítem se describe como: © 2013 Peña-Chilet et al.; licensee BioMed Central Ltd.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.











